KG2
Kaplan-Meier Graphs by Treatment Arm
KG
The same setup as in KG1 is used.
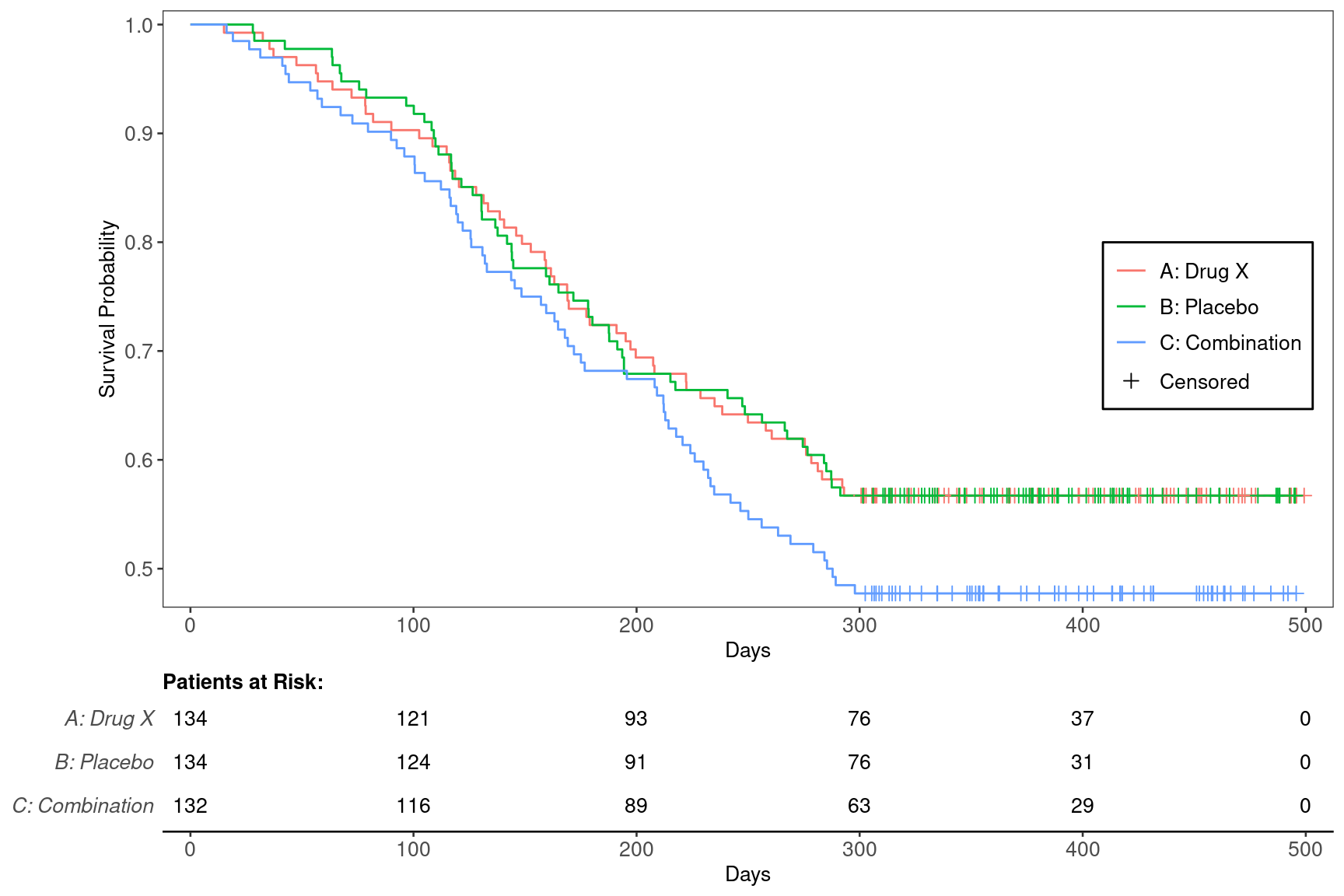
We can produce the basic plot using the g_km() function from tern.
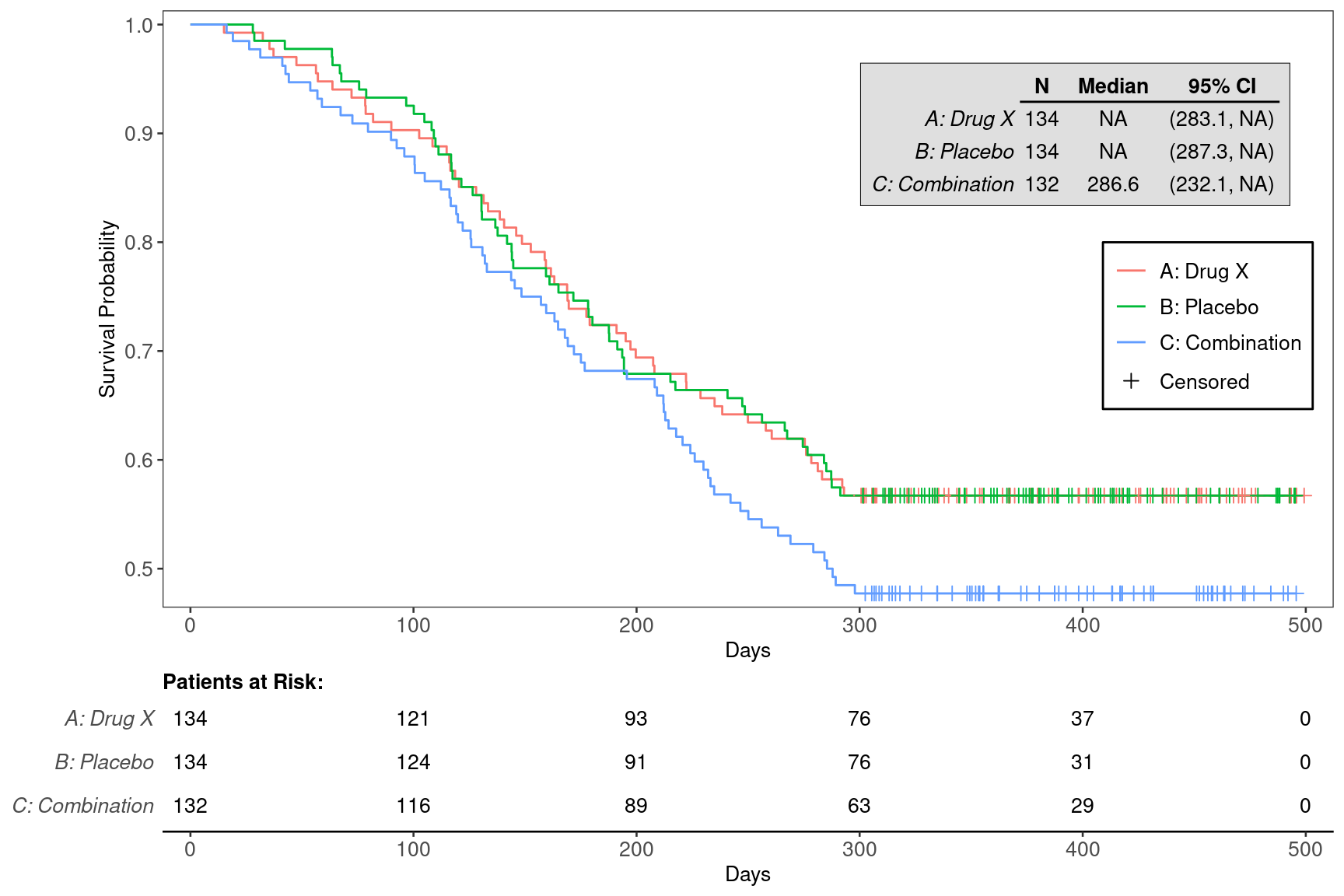
We can also choose to annotate the plot with the median survival time for each of the treatment arms using the annot_surv_med = TRUE option.
R version 4.5.2 (2025-10-31)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] dplyr_1.2.1 tern_0.9.10 rtables_0.6.15 magrittr_2.0.5
[5] formatters_0.5.12
loaded via a namespace (and not attached):
[1] generics_0.1.4 tidyr_1.3.2 stringi_1.8.7
[4] lattice_0.22-9 digest_0.6.39 evaluate_1.0.5
[7] grid_4.5.2 RColorBrewer_1.1-3 fastmap_1.2.0
[10] jsonlite_2.0.0 Matrix_1.7-5 backports_1.5.1
[13] survival_3.8-6 purrr_1.2.1 scales_1.4.0
[16] codetools_0.2-20 Rdpack_2.6.6 cli_3.6.5
[19] nestcolor_0.1.3 rlang_1.2.0 rbibutils_2.4.1
[22] cowplot_1.2.0 splines_4.5.2 withr_3.0.2
[25] yaml_2.3.12 otel_0.2.0 tools_4.5.2
[28] checkmate_2.3.4 ggplot2_4.0.2 forcats_1.0.1
[31] broom_1.0.12 vctrs_0.7.2 R6_2.6.1
[34] lifecycle_1.0.5 stringr_1.6.0 htmlwidgets_1.6.4
[37] pkgconfig_2.0.3 pillar_1.11.1 gtable_0.3.6
[40] glue_1.8.0 xfun_0.57 tibble_3.3.1
[43] tidyselect_1.2.1 knitr_1.51 dichromat_2.0-0.1
[46] farver_2.1.2 htmltools_0.5.9 rmarkdown_2.31
[49] labeling_0.4.3 random.cdisc.data_0.3.16 compiler_4.5.2
[52] S7_0.2.1 Reuse
Copyright 2023, Hoffmann-La Roche Ltd.